Comparative Genomic Hybridization (CGH) is a technique based on microarray technology, in which it is possible to identify and analyze the numerical and/or structural chromosomal alterations of the embryo in a single test.
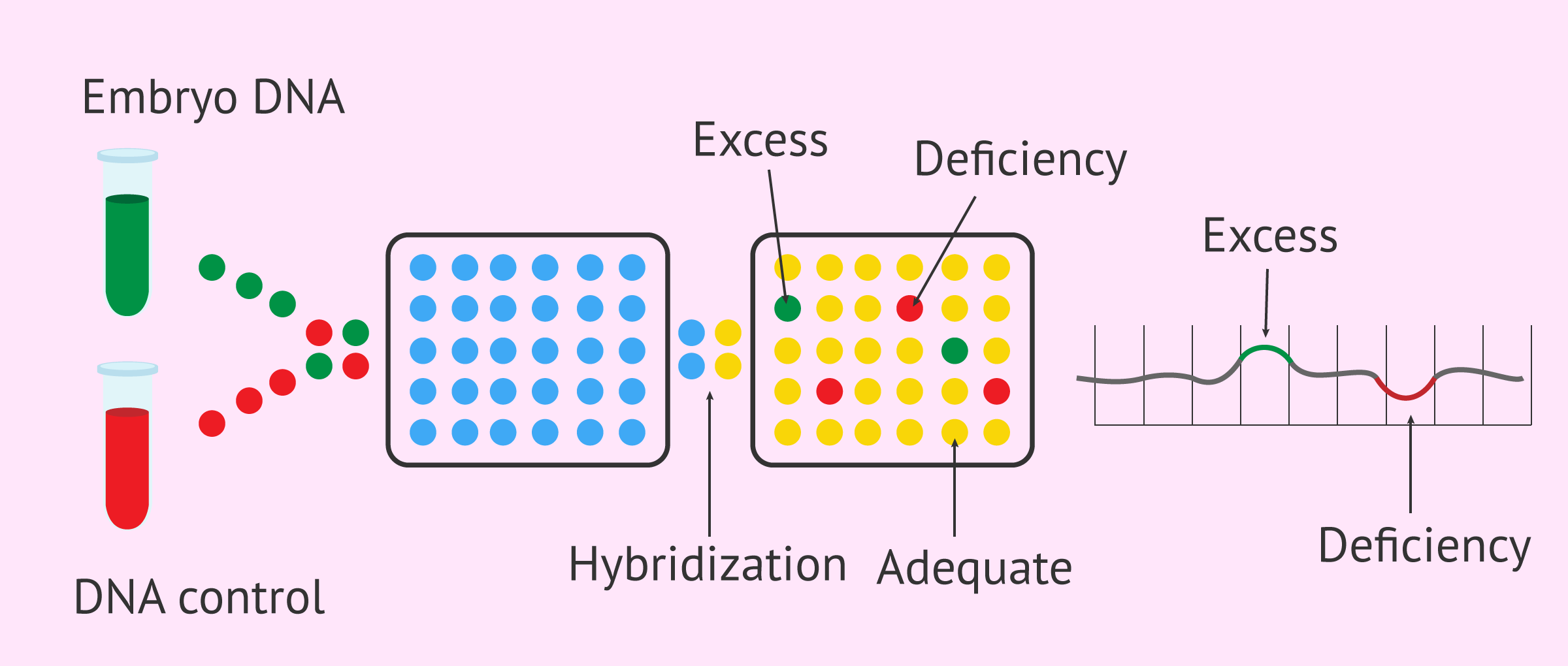
For this purpose, a sample of the embryonic DNA (marked with a green fluorescent probe) is compared with a sample of the control DNA (marked with a red fluorescent probe) to see if there is any deviation in the embryo from the reference sample. If the DNA from both samples matches, yellow fluorescence is observed in the array plate, resulting from the combination of green and red fluorescent molecules. On the other hand, if genetic information is missing from the embryo's DNA, red fluorescence will be observed. If, on the other hand, the embryonic DNA is duplicated, that is, it is found in a greater quantity than normal, the fluorescence emitted will be green.
The information obtained in the array is processed with bioinformatic tools to be able to interpret it by means of a graph. In it, the DNA regions in which the genetic content is adequate (yellow) will be seen in the central part, while the DNA losses (red) will be seen displaced downwards and the gains (green) displaced upwards. Since the location and order of the regions of the DNA contained in the different chromosomes is known, it is possible to know which regions of the DNA are affected by the deletions and duplications in the analyzed chromosome.
Great post, thanks for sharing! Love your pics.